Assay for reading six molecular pathways simultaneously developed
Posted: 16 December 2019 | Victoria Rees (Drug Target Review) | No comments yet
Researchers have developed a luciferase assay that can be used to monitor the effects of treatments on six different molecular targets at once.


Researchers have developed a novel approach, expanding the number of molecular pathways that can be studied on a cell sample with a dual luciferase assay from two to six.
The research was conducted at the Baylor College of Medicine, US and demonstrates that multiplexed hextuple luciferase assaying is possible.
According to the team, the technique can also be used to monitor the effects of experimental treatments on multiple molecular targets acting within these pathways.
The researchers say that the new assay is sensitive, saves time and expenses when compared to traditional approaches. It also reduces experimental error and can be adapted to any research field that uses a dual luciferase assay.
“One of the interests of our lab is to have a better understanding of the processes involved in cancer. Cancer usually originates through changes on many different genes and pathways, not just one, and currently, most cell-based screening assays conduct single measurements,” said corresponding author Dr Koen Venken, Assistant Professor of Biochemistry and Molecular Biology and Pharmacology and Chemical Biology at Baylor.
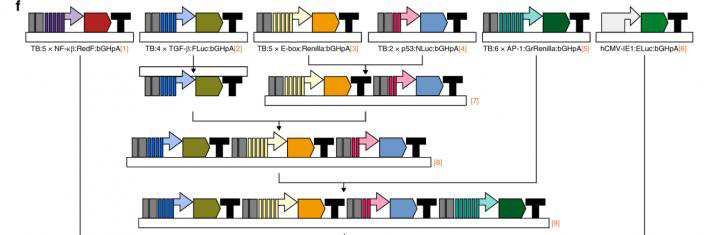

Synthetic assembly cloning for inserting multiple luciferase reporters into a single vector (credit: Nature Communications/The Venken lab).
The team focused on developing a multiplexed method to detect many signals from complex systems, such as living cells. They produced an assay that included six luciferases, each one emitting a unique bioluminescence that could be distinguished from the others, to reveal the activity of a particular pathway.
“To engineer and deliver the luciferase system to cells, we used a ‘molecular Lego’ approach,” said co-author Dr Lyra Chang, post-doctoral researcher at the Center for Drug Discovery at Baylor. “This consists of connecting the DNA fragments encoding all the biological and technological information necessary to express each luciferase gene together sequentially forming a single DNA chain called vector. This single vector enters the cells where each luciferase enzyme is produced separately.”
…the technique can also be used to monitor the effects of experimental treatments on multiple molecular targets”
Treating the cells with a single multi-luciferase gene vector instead of using six individual vectors decreased the variability between biological replicates and provided an additional level of experimental control. This allowed for simultaneous readout of the activity of five different pathways, compared to just one using traditional approaches, providing a much deeper understanding of cellular pathways.
“In addition to applications in cancer research, as we have shown in this work, our multiplex luciferase assay can be used to study other cellular pathways or complex diseases across different research fields,” Venken said. “For instance, the assay can be adapted to study the effect of drugs on insulin sensitivity in different cells types, the immune response to viral infections or any other combinations of pathways.”
The study was published in Nature Communications.
Related topics
Assays, Drug Targets, Hit-to-Lead, Molecular Targets, Research & Development, Screening
Related organisations
Baylor College of Medicine
Related people
Dr Koen Venken, Dr Lyra Chang