Antibodies developed from llama show promise in fighting COVID-19
Posted: 30 April 2020 | Victoria Rees (Drug Target Review) | No comments yet
Engineering antibodies found in a llama, researchers have developed a potential COVID-19 therapy which they say can neutralise SARS-CoV-2.

A group of researchers has identified a potential effective COVID-19 treatment, developed from antibodies found in a llama. The research was conducted by scientists at the University of Texas (UT) at Austin, the National Institutes of Health (NIH), both US, and Ghent University, Belgium.
When llamas’ immune systems detect foreign invaders such as bacteria and viruses, these animals (and other camelids such as alpacas) produce two types of antibodies: one that is similar to human antibodies and another that is about a quarter of the size. These smaller ones, called single-domain antibodies or nanobodies, can be nebulised and used in an inhaler.
“That makes them potentially really interesting as a drug for a respiratory pathogen because you’re delivering it right to the site of infection,” said Daniel Wrapp, a graduate student at UT Austin and co-first author of the paper.


Scientists were inspired by antibodies produced by this llama, named Winter, to develop their antibody against SARS-CoV-2. Winter is four years old and still living on a farm in the Belgian countryside operated by Ghent University’s Vlaams Institute for Biotechnology [credit: Tim Coppens].
In 2016, the researchers were studying two earlier coronaviruses: SARS-CoV-1 and MERS-CoV. In a process similar to human immunisation, a particular llama was injected with stabilised Spike (S) proteins from those viruses over the course of about six weeks. The S protein is found on the coronavirus that causes COVID-19 and allows the virus to break into host cells.
Next, the researchers collected a blood sample and isolated antibodies that bound to each version of the S protein. One showed promise in stopping a virus that displays S proteins from SARS-CoV-1 from infecting cells in culture.
The first antibodies the team identified in the initial SARS-CoV-1 and MERS-CoV tests included one called VHH-72, which bound tightly to S proteins on SARS-CoV-1. This prevented a pseudotyped virus (a virus that cannot make people sick and has been genetically engineered to display copies of the SARS-CoV-1 S protein on its surface) from infecting cells.
When SARS-CoV-2 emerged and triggered the COVID-19 pandemic, the team wondered whether the antibody they discovered for SARS-CoV-1 would also be effective against its viral cousin. They discovered that it did bind to SARS-CoV-2’s S protein, albeit weakly. They engineered it to bind more effectively by linking two copies of VHH-72, which they then demonstrated neutralises a pseudotyped virus sporting S proteins from SARS-CoV-2. According to the researchers, this is the first known antibody that neutralises both SARS-CoV-1 and SARS-CoV-2.
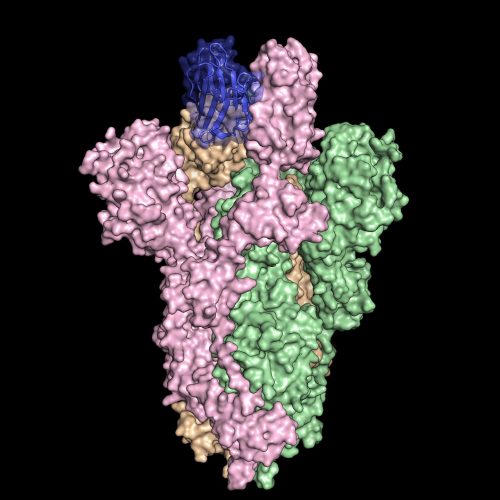

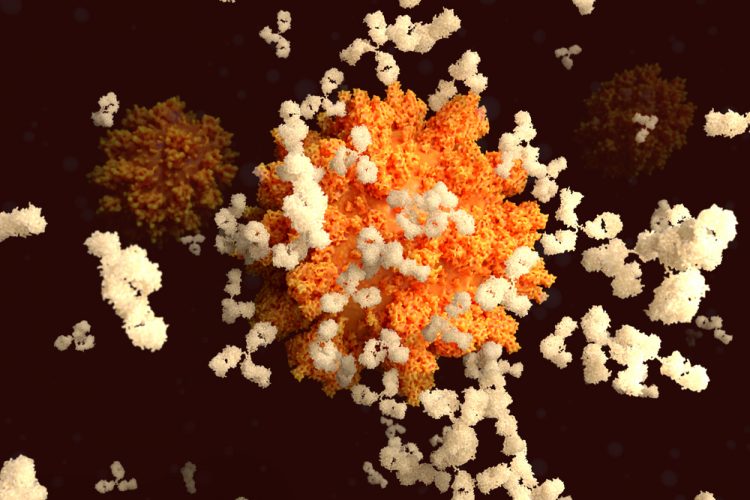
Inspired by a special kind of antibody produced by llamas, researchers created an antibody dubbed VHH-72Fc (blue) that binds tightly to the spike protein on SARS-CoV-2 (pink, green and orange), blocking the virus from infecting cells in culture. The spike protein structure was discovered by part of the same research team and published in the journal Science on February 19, 2020 [credit:
University of Texas at Austin].
“This is one of the first antibodies known to neutralise SARS-CoV-2,” said Jason McLellan, associate professor of molecular biosciences at UT Austin and co-senior author..
The team is now preparing to conduct pre-clinical studies in animals such as hamsters or nonhuman primates, with the hopes of subsequently testing in humans.
“Vaccines have to be given a month or two before infection to provide protection,” McLellan said. “With antibody therapies, you’re directly giving somebody the protective antibodies and so, immediately after treatment, they should be protected. The antibodies could also be used to treat somebody who is already sick to lessen the severity of the disease.”
The team will report their findings on 5 May in the journal Cell. The paper is currently available online as a “pre-proof,” meaning it is peer-reviewed but undergoing final formatting.
Related topics
Antibodies, Antibody Discovery, Disease Research, Drug Targets, Immunology, Proteomics
Related conditions
Coronavirus, Covid-19, Middle East Respiratory Syndrome (MERS), Severe Acute Respiratory Syndrome (SARS)
Related organisations
Ghent University, National Institutes of Health (NIH), University of Texas at Austin
Related people
Daniel Wrapp, Jason McLellan