Mapping SARS-CoV-2 infection in the airway
Posted: 2 June 2020 | Hannah Balfour (Drug Target Review) | No comments yet
Researchers found there is a gradient of SARS-CoV-2 infectivity down the airway and that the severe pneumonia symptoms may be caused by aspiration of oral contents into the lungs.
Scientists have characterised how SARS-CoV-2, the coronavirus causing the COVID-19 pandemic, effectively infects the nasal cavity, but infects and replicates progressively less well in cells lower down the respiratory tract, including the lungs.
The study published in Cell, suggests the virus becomes established in the nasal cavity first, before being aspirated to the lungs where it causes the more severe symptoms including pneumonia. This later stage only occurs in some cases.
“If the nose is the dominant initial site from which lung infections are seeded, then the widespread use of masks to protect the nasal passages, as well as any therapeutic strategies that reduce virus in the nose, such as nasal irrigation or antiviral nasal sprays, could be beneficial,” said study co-senior author Dr Richard Boucher, the James C. Moeser Eminent Distinguished Professor of Medicine and Director of the Marsico Lung Institute at the University of North Carolina (UNC) School of Medicine, US.
“This is a landmark study that reveals new and unexpected insights into the mechanisms that regulate disease progression and severity following SARS-CoV-2 infection,” said Dr Ralph Baric, co-senior author of the study, the William R. Kenan Distinguished Professor of Epidemiology at the UNC Gillings School of Public Health who also holds a microbiology faculty appointment at the UNC School of Medicine. “In addition, we describe a new reverse genetic platform for SARS-CoV-2 allowing us to produce key indicator viruses that will support national vaccine efforts designed to control the spread and severity of this terrible disease.”
The team sought to understand which cells in the airway are infected by SARS-CoV-2 and how it gets into the lungs in the patients who develop pneumonia.
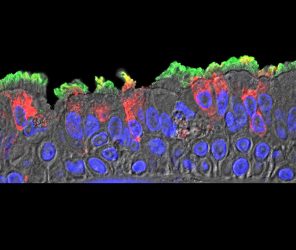
SARS-CoV-2 (red) infected ciliated cells in the COVID-19 patient’s bronchi [credit: Takanori Asakura, PhD, UNC School of Medicine].
In one round of experiments, the scientists used different isolates of SARS-CoV-2 to see how efficiently they could infect cultured cells from different parts of the human airway. They observed a gradient from relatively high infectivity in cells lining the nasal passages, to less infectivity in cells lining the throat and bronchia, to relatively low infectivity in lung cells. Correspondingly the angiotensin-converting enzyme 2 (ACE2), which SARS-CoV-2 is known to use to invade cells, was more abundant on nasal-lining cells and less abundant on lower airway cells. The expression pattern of ACE2 receptors could potentially account for the gradient of infectivity, according to the team.
Their other experiments focused on two protein-cleaving enzymes found on human cells, TMPRSS2 and furin, which are thought to be used by SARS-CoV-2 to cleave and re-shape viral proteins so they can enter human cells. The team observed that when these human enzymes are more abundant, the coronavirus had an increased ability to infect cells and make copies of itself.
The cells the researchers found SARS-CoV2 to infect were airway epithelial cells and – to a lesser extent – pneumocyte lung cells, which help transfer inhaled oxygen into the bloodstream. However, the virus did not infect any other airway cells, including club cells which express both ACE2 and TMPRSS2. Moreover, the same types of airway epithelial cells from different human donors, especially lower-airway epithelial cells, tended to vary significantly in their susceptibility to infection. Suggesting there are other underlying factors that determine infectivity, yet to be discovered.
The team also mapped the sites of SARS-CoV-2 infection in the lungs of several patients who had died from COVID-19. According to the researchers, these sites were patchy and had characteristics consistent with the hypothesis that these sites had originated from infection higher in the airway.
The hypothesis that aspiration of oral contents into the lung is a significant contributor to COVID-19 pneumonia is also consistent with the observations that people at higher risk for severe lung disease – the elderly, obese and diabetic – are more prone to aspiration, especially at night said the team.
They also tested whether neutralising antibodies for related coronaviruses, Severe acute respiratory syndrome (SARS) and Middle eastern respiratory syndrome (MERS), offered protection against SARS-CoV-2 infection. The other than blood serum from two of the five SARS samples, antibodies showed little ability to neutralise SARS-CoV-2 infectivity in cultured cells.
“These results, using some novel and innovative methodology, open new directions for future studies on SARS-C0V-2 that may guide therapeutic development and practices for reducing transmission and severity of COVID-19,” said James Kiley, Director of the Division of Lung Diseases at the National Heart, Lung, and Blood Institute, part of the National Institutes of Health (NIH).
The team highlighted that aside from their findings, the study also involved the development of key laboratory tools, including a version of SARS-CoV-2 re-engineered to carry a fluorescent beacon, that they state should be useful in future research.
Related topics
Antibodies, Disease Research, Drug Targets, Enzymes, Immunology, Protein, Protein Expression, Proteomics
Related conditions
Coronavirus, Covid-19
Related organisations
National Heart Lung and Blood Institute (NHLBI), University of North Carolina (UNC) Gillings School of Public Health, University of North Carolina (UNC) School of Medicine, US National Institutes of Health (NIH)
Related people
Dr Ralph Baric, Dr Richard Boucher, James Kiley