Study finds activation sequence of COVID-19 S protein is cleaved by furin protease
Posted: 6 May 2020 | Victoria Rees (Drug Target Review) | No comments yet
Researchers have shown that the activation sequence the SARS-CoV-2 S protein is cleaved by the cellular enzyme furin which is also required for the infection of lung cells.
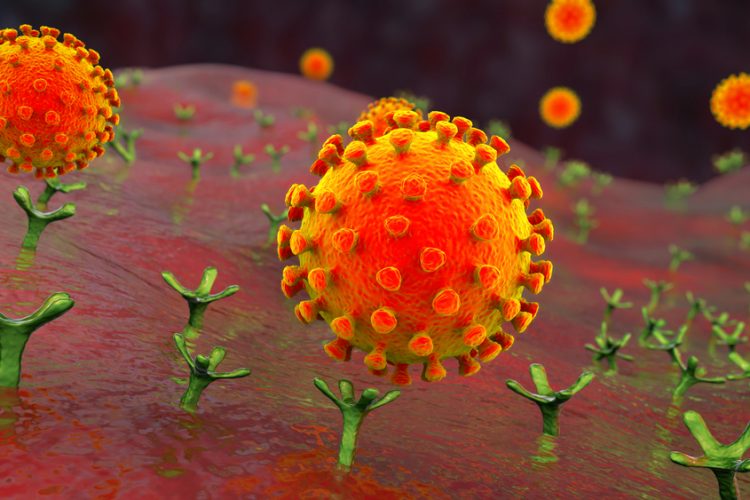
A group of researchers has shown that the activation sequence of the Spike (S) protein of SARS-CoV-2, which causes COVID-19, is cleaved by the cellular enzyme furin. They also found that this cleavage is important for the infection of lung cells.
The study was conducted at the Infection Biology Unit of the German Primate Center (DPZ) – Leibniz Institute for Primate Research. According to the researchers, these results define new starting points for therapy and vaccine research.
The S protein on the SARS-CoV-2 surface serves as a key for the virus to enter host cells. It facilitates viral attachment to cells and fuses the viral and cellular membranes, thereby allowing the virus to deliver its genome into the cell, which is essential for viral replication. For this, activation sequences of the S protein need to be cleaved by cellular enzymes, called proteases. The S protein carries an activation sequence at the so-called S1/S2 cleavage site, which is similar to those observed in highly pathogenic avian influenza viruses, but which has so far not been found in viruses closely related to SARS-CoV-2. The importance of this sequence for the virus was so far unknown.
In their current study, the infection biologists were able to show that the S1/S2 activation sequence of the S protein is cleaved by the cellular protease furin and that this cleavage event is essential for the infection of lung cells. It is also important for the fusion of infected cells with non-infected cells, which might allow the virus to spread in the body without leaving the host cell.
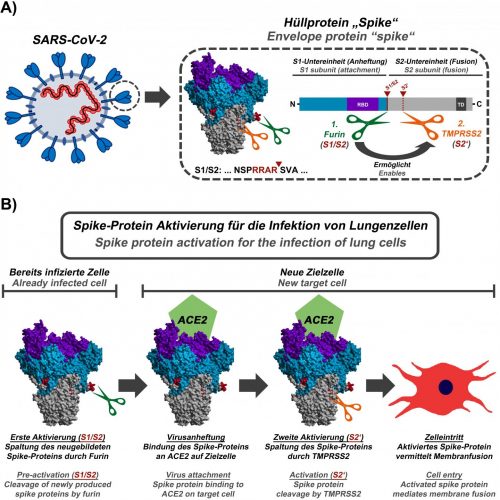
(A) Schematic representation of SARS-CoV-2, the viral S protein and the cleavage sites for furin (green, S1/S2 position; the cleavage sequence is shown below the protein structure) and TMPRSS2 (orange, S2′ position). (B) First, in already infected cells, the enzyme furin cuts the S protein at the S1/S2 site. The S protein then mediates viral attachment to a new host cell. In order to efficiently enter the cell, the S protein still needs to be activated by the enzyme TMPRSS2. Activation by TMPRSS2 is only possible if the S protein has previously been cleaved by furin [credit: Markus Hoffmann].
“Our results suggest that inhibition of furin should block the spread of SARS-CoV-2 in the lung,” said one of the lead researchers Stefan Pöhlmann, head of the Infection Biology Unit at DPZ. “Furthermore, our present study and previous work demonstrate that the virus uses a two-step activation mechanism: In infected cells, the S protein has to be cleaved by the protease furin so that newly formed viruses can then use the protease TMPRSS2 for further cleavage of the spike protein, which is important for the entry into lung cells.”
For a live attenuated vaccine to trigger a strong immune response, the researchers say it has to be able to replicate in the body to a limited extent, for example locally at the site of injection. “SARS-CoV-2 variants, in which the activation sequence for furin has been removed, could be used as a basis for the development of such live attenuated vaccines, since the lack of cleavage of the S protein should greatly limit the spread of the virus in the body. A sufficiently attenuated virus would no longer be able to cause disease, but would still enable the immune system to react to the pathogen and for example, produce neutralising antibodies,” said one of the lead researchers Markus Hoffmann, also first author of the study.
The study was published in Molecular Cell.
Related topics
Disease Research, Drug Targets, Protein, Proteomics, Research & Development
Related conditions
Coronavirus, Covid-19
Related organisations
German Primate Center (DPZ)
Related people
Markus Hoffmann, Stefan Pöhlmann