Mutations in SARS-CoV-2 Spike could lead to greater infectivity, study shows
Posted: 16 February 2021 | Victoria Rees (Drug Target Review) | No comments yet
Comparing the original SARS-CoV-2 Spike protein with a mutated version, researchers have potentially revealed why the mutated version is dominant.
Researchers have compared the original SARS-CoV-2 Spike (S) protein with a mutated version which arose last spring. They have found structural differences which could help to explain why the mutated version remains the dominant form circulating in all recent variants of concern, such as the UK and South African strains. The research was conducted at the Francis Crick Institute, UK.
As SARS-CoV-2 has spread and evolved, mutations impacting the structure of the viral S protein have affected how it binds to and infects cells, which has had huge implications for the spread of disease as new variants dominate in part due to changes in the fitness of the virus.
The researchers used cryo-electron microscopy (cryo-EM) to gain detailed images of a variant form of the SARS-CoV-2 S protein. They then compared this to a previous study examining the original ‘Wuhan’ form of the virus. The variant S protein called G614 is a mutated version which first arose around spring 2020 and is now the dominant version of the S protein in circulation, including in the recently identified variants of concern.
By analysing their structures, the researchers found that the mutated G614 S protein would more easily bind to human cells due to its open and flexible structure. While the original S protein adopted a generally more closed form, they observed that the mutant protein could adopt a greater number of open and erect conformations, which would prime the protein for binding to the human cell receptor angiotensin-converting enzyme 2 (ACE2).
“As this mutant version likely binds more easily to cells, it has an advantage over the original S protein. This could be one reason why this variant now predominates over the initial protein and it is more wide-spread in circulation,” said co-lead author of the study Donald Benton.
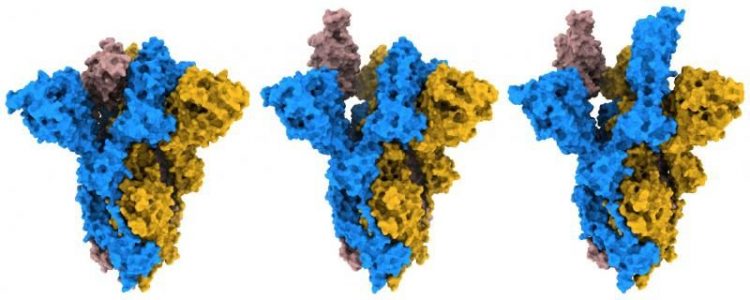
Different conformations observed of D614G SARS-CoV-2 variant S protein [credit: Francis Crick Institute].
However, the researchers suggest that this advantage could eventually become problematic for the virus. This is because its more open and flexible structure could mean that it is more exposed to circulating antibodies.
Antoni Wrobel, co-lead author of the study, said: “During the first wave of infection, most people did not have antibodies to SARS-CoV-2, so this open-structured mutant S protein could have been beneficial to the virus. However, as time goes on, more people will have antibodies as a result of previous infection or vaccination. And as this mutant S protein presents more surface area, it is more exposed to these antibodies, which could be a disadvantage.”
The study was published in PNAS.
Related topics
Disease Research, Imaging, Microscopy, Protein, Proteomics
Related conditions
Covid-19
Related organisations
Francis Crick Institute
Related people
Antoni Wrobel, Donald Benton