From a database of more than 200,000 high-resolution, three-dimensional images of human induced pluripotent stem cells, researchers have devised a model to quantify cell shape and internal organization. Susanne Rafelski, Deputy Director of the Allen Institute for Cell Science, revealed details of their study to Drug Target Review.

A recently published study1 from a large team of cell biologists sheds new light on cellular organisation and variability. Researchers at the Allen Institute for Cell Science, a division of the Allen Institute in Seattle, US, developed a unique mathematical framework to uncover internal organisation of cellular structures in human induced pluripotent stem cells (hiPSCs) while taking account of the wide range of variation that naturally occurs in cell shapes.
The team had to first develop standardised methods to create and image fluorescently tagged hiPSC cell lines and to segment the tagged internal structures, and the cells, before they were able to map internal organisation. All of this was built for three-dimensional (3D) images, posing extra challenges to the team as most existing methods were developed for two-dimensional (2D) images.
Dr Susanne Rafelski, who led the study, spoke with Drug Target Review about how the study was conducted, how it might apply to diseases, and future directions.
Can you give a brief overview of your research methods?
This project started in the early days of the Allen Institute for Cell Science, nearly seven years ago. We first figured out how to tag structures inside stem cells with fluorescent proteins while keeping the cells as healthy as possible. The result was the Allen Cell Collection2 of gene-edited hiPSC lines, 25 of which were used in this study. Next, we needed to figure out the best way to image these cells live and in 3D so that we could view structures in high resolution without killing them. Finally, we faced the daunting task of developing computational image analysis tools to transform our 3D microscopy images into segmented images of individual cells and their structures. At that point, we had our dataset in hand and began developing the generalisable analysis framework that was the basis of our recent paper. This framework is based on two coordinate systems for understanding the variance and organisation of hiPSCs: one for cell shape and one to map the interiors of the cells.

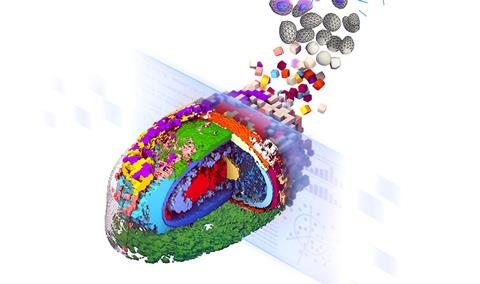
Stylised representation of an integrated average morphed cell showing the location of 17 select cellular structures (Credit: Thao Do, Allen Institute for Cell Science)[/caption]
Our study encompasses all the expertise of our diverse team at the Allen Institute for Cell Science and is truly a collaborative effort. Our 84 co-authors include molecular biologist experts in gene editing, cell biologist experts in microscopy imaging, computational scientist experts in image and data analysis, software engineer experts in database infrastructure, and visualisation experts to develop tools for us all to explore the data. All these researchers effectively combined their expertise for this seminal work.
What is the current understanding of integrated intracellular organisation in hiPSCs, and how does it differ from other cell types?
This was a big unknown when we started this project, for hiPSCs and really any kind of cell. People had studied relationships between some individual structures in cells, such as the position of the Golgi apparatus in migrating cells, but overall internal integrated cellular organisation was not known - not in the way we needed in order to measure and quantify it. When we started tagging structures and looking at them in our hiPSC lines, the images were brand new because people just weren’t doing high-resolution 3D imaging of organisation in hiPSCs.
Our study not only builds a framework for stem cell organisation for the benefit of future experiments with this particular cell type, but also provides a roadmap for researchers to do similar analyses in other kinds of cells.
Why is understanding cell organisation so important to disease research?
We know cell organisation matters — pathologists have used visual inspection of the arrangement of cell components for decades to diagnose progression and prognosis of diseases such as cancer. But we haven’t tapped into all the information that lies in cell organisation in a quantitative way. In disease, what goes wrong in cells can often be seen by eye. Cancer pathology is a clear example of that. But by the time you can spot a disease under the microscope, it’s a big sledgehammer. What we can see by numbers and distributions of populations is far more subtle. You don’t want to catch disease when it’s a sledgehammer, you want to catch it when smaller changes are happening, because disease is the additive effect of small changes until you get to big changes.
To catch small changes, you have to have rigorous, statistical means of identifying them. That’s what we’re hoping this work can lead to.
Can you tell us more about WTC-11 hiPSC Single-Cell Image Dataset v1?
This dataset3 is so exciting, because there are very few high-resolution, 3D single-cell image datasets out there that cover such a wide range of structures. That creates a resource for scientists, including data science researchers who want to apply machine learning to cell images; now they have more than 200,000 cells they can play with. It also allows scientists to explore hypotheses about the cell biology of undifferentiated stem cells. A lot of other screening datasets use 2D images and cancer cell lines, but we needed three dimensions to analyse hiPSCs. We needed a different type of imaging, a different kind of approach. We know there is a lot of interest in utilising hiPSCs for both basic and disease-focused research, and that’s why we have made them available for anyone in the community- both non-profit and commercial research- to use.
What challenges did you encounter in your research?
There were challenges in each step along the way. We first needed to figure out how to tag structures in cells while keeping them as healthy and normal as possible. Overcoming those challenges enabled us to create the Allen Cell Collection. The next challenge was to image the cells in a standardised way. We added in some automation and developed very particular ways to image these cells, the details of which were published in the Nature paper and a protocol paper we are currently working on. Once we had the images, we needed to establish how to segment them — how to define the boundaries of each structure inside the cell and how to define the cell edges. That was a big challenge that led to the development of a software toolkit, the Allen Cell & Structure Segmenter,4 which is also openly available. Finally, there was the general challenge of integrating and quantifying all our data, which is the framework we describe in the paper.
Can you discuss any notable variations in integrated intracellular organisation that you observed in iPSCs, and what implications these variations have for our understanding of cellular function?
We developed our metrics to look at the mean location and variation in individual cellular structures, and in pairs of structures in the cell. When viewing healthy hiPSCs in interphase, we found that the wiring of internal organisation is highly consistent across cells, far more so than we expected. We then looked at two different conditions: cells at the edges of colonies, and cells undergoing mitosis. In edge cells, the wiring diagram is still consistent, but the location of many key structures inside the cell changes. That’s likely due to these edge cells not being surrounded by cells on all sides. What this tells us is that the location of cellular structures can change without changing the variability in those locations or the relationships between them. We didn’t know that before. For cells in mitosis, all of our measurements of location and wiring change, but the specific types of changes and their timing during mitosis depend on the specific structure. In all of these instances, we found that changes in location of an individual structure precede changes in their wiring, and that changes in individual structure variations generally precede changes in pairwise interactions, with a few key exceptions. This hints at a possible hierarchy of dependencies as cells reorganise, which we look forward to exploring further together with the community.
How might your research on integrated intracellular organisation in iPSCs contribute to our understanding of diseases and potential treatments?
One important aspect of our paper was that we needed to build the framework to capture the full spectrum of normal cell shapes in order to map the interiors of the cells in abnormal conditions. Because hiPSCs, and generally any population of cells, have such a wide range of shape, we needed to first cluster similarly shaped cells together so that we could compare internal organisation in a controlled manner. That means that, for researchers working on a specific disease or a drug treatment, provided those perturbations don’t drastically change the shape of the cells, they could use the same framework to compare internal organisation between healthy hiPSCs and their perturbed cells, to see if internal phenotypes have changed. Our frameworks are all quantitative so we can put numbers on all these differences, both the mean and the variation for the perturbed condition.
What future directions do you see for research in this area, and what new technologies or techniques might help advance our understanding of intracellular organisation in iPSCs?
Now that the framework has been established, and in keeping with the Allen Institute’s dedication to open science, we’re hoping scientists from around the world will use our method to study their own cell lines and uncover foundational principles of cell behaviour and organisation. Uncovering the rules of a “normal cell” and learning the full variation of what “normal” looks like is key to deepening our understanding and helping us find better treatments for disease. We and others are already applying some of the concepts in this paper to analysing cell organisation in other cell types and states, including disease states, and extending the concepts to study not just the structures inside the cell but also to look for patterns in the locations of proteins making up those structures.
Uncovering the rules of a “normal cell” and learning the full variation of what “normal” looks like is key to deepening our understanding and helping us find better treatments for disease.
Any other comments?
It was truly a privilege to be a part of such a large and interdisciplinary team effort to create something that we hope will be a great resource for the community. We anticipate that it will fill in significant gaps to be able to measure, analyse and understand how cells are organised and how this impacts cell behaviour and function.
Author bios:
 Dr Susanne Rafelski
Dr Susanne Rafelski
Susanne is the Deputy Director of the Allen Institute for Cell Science, which aims to understand the principles by which human induced pluripotent stem cells (hiPSC) establish and maintain robust dynamic localisation of cellular structures, and how cells transition between states during differentiation and disease. Her life-long scientific goal is to decipher the patterns and rules that transform the overwhelming complexity found inside cells into functioning units of life.

Dr Rachel Tompa
Rachel is Senior Editor at the Allen Institute. A former molecular biologist, she’s been writing about health and science for more than 15 years, covering a wide variety of topics including the microbiome, pregnancy, global health, science policy and infectious disease. She is also co-host and co-producer of the Allen Institute’s podcast, Lab Notes.
References:
- Viana MP, Chen J, Knijnenburg TA, et al. Integrated intracellular organisation and its variations in human IPS cells. Nature. 2023;613(7943):345–54.
- Cell line catalog [Internet]. ALLEN CELL EXPLORER. [cited 2023Apr4]. Available from: https://www.allencell.org/cell-catalog.html
- Cell feature explorer [Internet]. Cell Feature Explorer. [cited 2023Apr4]. Available from: https://cfe.allencell.org/
- Allen Cell & Structure segmenter [Internet]. ALLEN CELL EXPLORER. [cited 2023Apr4]. Available from: https://www.allencell.org/segmenter.html