In an exclusive interview, Dr Neville Sanjana, Associate Professor of biology at NYU and a core faculty member at the New York Genome Center, discusses the breakthrough study on STING-seq.

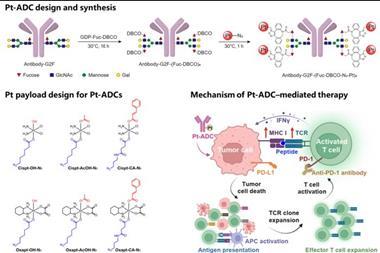
Co-led by Dr Sanjana, the research study combines CRISPR gene editing and single-cell sequencing to identify causal genetic variants and enhance our understanding of the noncoding genome. Published in Science, the study offers new possibilities for identifying therapeutic targets in genetically influenced diseases. The workflow, which has been termed STING-seq, integrates biobank-scale GWAS data with CRISPR and single-cell sequencing techniques, allowing for the identification of target genes associated with noncoding variants for blood traits. This method has the potential to advance precision medicine and targeted therapies across various diseases and traits. Dr Sanjana elaborated on the study for Drug Target Review.
What are the main challenges in human genetics research when it comes to understanding which parts of the genome drive specific traits or contribute to disease risk?
Most human genetic variation is found outside of genes that code for proteins. This presents two major challenges. First, it is challenging to understand which variants are causal and which are simply in linkage with the causal variant. Put simply, correlation is not causation. By perturbing specific variants using CRISPR, we can test which ones alter gene expression or disease-associated phenotypes. The second challenge is that for noncoding variants, even if we know the causal variant, we often do not know what the target gene is; that is, how the causal variant impacts diseases or traits.
How can combining genetic association studies, gene editing, and single-cell sequencing help to address these challenges and discover causal variants and genetic mechanisms for blood cell traits?
Indeed STING-seq — which combines genome-wide association studies (GWAS), gene editing and single-cell sequencing — lets us pinpoint causal variants and their target genes and does so at scale. STING-seq lets us interrogate hundreds or thousands of genomic regions to pinpoint those that impact gene expression and tells us which genes change.
How does STING-seq address the challenge of directly connecting genetic variants to human traits and health, and how can it help identify drug targets for diseases with a genetic basis?
By identifying target genes for noncoding variants that are either protective or associated with disease, we have powerful genetic evidence that these target genes are worth pursuing as drug targets for the disease. This approach is quite general and can work for any disease where we have variants from genetic association studies.
What was the scientific breakthrough in the treatment of sickle cell anaemia, and how did combining GWAS with CRISPR lead to the identification of causal variants and innovative therapies?
This is a great question. Most people carrying two copies of the sickle mutation in the adult haemoglobin gene will get sickle-cell anaemia. Yet some people do not. These people seem to be expressing a different form of haemoglobin, called foetal haemoglobin. Normally, we do not express foetal haemoglobin after birth but some people have a persistent expression of this protein. For those with persistent expression, genetic association studies identified a large region of the genome near a gene called BCL11A. This is a repressor of foetal haemoglobin. If you turn off BCL11A, you reactivate foetal haemoglobin.
In collaboration with Dan Bauer, Stu Orkin and Matt Canver, we performed a CRISPR screen about a decade ago over this region of the genome and identified a noncoding region (a transcription factor binding site) that controls BCL11A expression. If you edit this site, you can turn off BCL11A and reactivate foetal haemoglobin. This CRISPR has now been used in about 100 patients and in nearly all cases, the reactivation of foetal haemoglobin seems to act as a durable cure for sickle-cell and other haemoglobin disorders.
How do you see the STING-seq approach being applied to other diseases beyond blood cell traits, and what are the challenges in scaling up this approach?
STING-seq can be used in any disease where we have some nominated variants and regions of the genome through genetic association studies. This includes not just blood traits but cardiovascular, autoimmune, neuropsychiatric and neurodegenerative disorders — to name a few broad categories. This encompasses many common diseases and we are optimistic that STING-seq will be a useful platform to rapidly understand which variants and genes are key players in these diseases.
The perennial challenge with STING-seq (and all cell-based assays) is finding the right cell type for the disease or trait. In some cases, a single cell type will probably suffice but, in many cases, it will be important to perform and compare STING-seq across several different cell types to understand how the cell context shapes the activity of these genetic variants.
What are the limitations of genome-wide association studies (GWAS) in identifying causal genes and variants, and how does STING-seq overcome these limitations?
The fundamental limitation of GWAS is that it measures correlations but does not yield a mechanistic, causative model. STING-seq connects human genetic variants identified through approaches like GWAS with the mechanistic basis for their activity (that is, target genes).
 Dr Neville Sanjana
Dr Neville Sanjana
Dr Neville Sanjana, PhD, is Associate Professor of biology at NYU, a core faculty member at New York Genome Center, and the study’s co-senior author.