Neuropilin-1: a key receptor in SARS-CoV-2 infection
Posted: 27 November 2020 | Nikki Withers (Drug Target Review) | No comments yet
Results of a recent study by researchers at the University of Bristol indicate neuropilin-1 is an important host factor for SARS-CoV-2 infection. The team suggests that blocking neuropilin-1 may be a valuable therapeutic intervention in the treatment of COVID-19. Nikki Withers spoke to one of the study’s lead investigators, Dr Yohei Yamauchi, to hear more about their findings.
ALTHOUGH PANDEMICS associated with coronaviruses (CoVs) have led to fatalities in the past, COVID-19 has taken the world by surprise. “The main difference between the past CoV outbreaks and this one is that SARS-CoV-2 is able to transmit between humans much more easily than the others,” explained Yohei Yamauchi, a virologist from the School of Cellular and Molecular Medicine at the University of Bristol in the UK.
His team, together with Peter Cullen and Boris Simonetti at the School of Biochemistry at Bristol, has potentially identified a mechanism that causes SARS-CoV-2 to be highly infectious and able to spread rapidly in human cells. Their findings, published in Science,1 describe how the virus’s ability to infect human cells can be reduced by inhibitors that block a newly discovered interaction between the virus and host, demonstrating a potential anti-viral treatment.
Increased transmissibility
When we discussed the differences between previous CoV outbreaks and the current pandemic, Yamauchi explained that all previous outbreaks have been relatively localised, although with higher fatality rates than SARS-CoV-2. Indeed, the first pandemic threat of a novel and deadly coronavirus was the virus causing Severe Acute Respiratory Syndrome (SARS-CoV) that emerged in late 2002. It was highly lethal (case fatality rate of 9.7 percent) but faded out after intense public health mitigation measures.2 Another coronavirus, the Middle East Respiratory Syndrome coronavirus (MERS-CoV), emerged in 2012 and had a very high case fatality rate – approximately 34 percent.2 Each virus caused around 800 deaths worldwide. It must be noted that MERS-CoV is widespread in dromedary camels and zoonotic cases continue to occur, unlike SARS-CoV, which emerged from wildlife and was eliminated from the intermediate host reservoir.2 However, with both these earlier viruses “unless there was intense travel between the region of the outbreak and another region of the globe, it was not spreading like it is right now,” said Yamauchi.
Although the new coronavirus is less deadly, it is far more transmissible than MERS-CoV or SARS-CoV. “It spreads like flu, but it causes more fatalities, more hospitalisations and requires more intensive care. We also do not have any vaccines against any of the coronaviruses but there are promising developments like the Pfizer/BioNTech and Moderna mRNA vaccines. Whether it confers long-term immunity, however, is unknown,” said Yamauchi. Furthermore, unlike seasonal coronaviruses, which cause common colds and mild respiratory symptoms, SARS-CoV-2 is not only highly infective and transmissive, but also readily infects organs outside of the respiratory system (such as the brain and heart) through mechanisms that are currently unclear. “COVID-19 patients are also presenting with other neurological symptoms, such as loss of their sense of taste or smell, or long COVID. This is slightly different to previous CoV pandemics.”
Receptor interactions
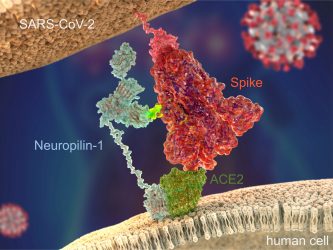
Figure 1: SARS-CoV-2 Spike protein (red) binds to the human cell surface receptor ACE2 (green), and
Neuropilin-1 (blue) via the Spike’s C-end Rule motif (fluorescent yellow) [credit: Peter Allen
and Ryan Allen, Second Bay Studios].
Since the start of the current outbreak, our understanding of the virus and how it infects cells has increased significantly. “In order to bind to human cells, it is known that the Spike protein of SARS-CoV-2 recognises and makes contact with a receptor called angiotensin-converting enzyme 2 (ACE2) on the surface, which triggers internalisation,” explained Yamauchi. However, in their recent study, the team discovered a secondary receptor that the virus recognises to cause infection, called neuropilin-1 (Figure 1). “We found that the presence of neuropilin-1 at the cell surface significantly enhanced the rate of infection and spread of the virus in human cells.”
Prior to their research, neuropilin-1 was already understood to have a multifaceted, regulatory function as a cell surface protein. “In 2009, there was a study from America3 showing that this neuropilin-1 cell surface protein can bind to a ligand, which has a specific amino acid sequence at the C-terminus called the C-end rule motif (CendR). Once it binds, the ligand triggers cellular events like vascular and tissue penetration, and internalisation of the ligand into cells,” explained Yamauchi. “The 2009 study had already spotted that this C-end rule peptide is prevalent in many pathogenic viruses such as avian influenza and Ebola. These viruses had acquired a multibasic cleavage site, which is processed by furin – an abundant protease in our bodies – liberating a C-end rule peptide.”
Our hypothesis was that the reason this virus spreads quite dramatically between humans could be due to viral Spike’s ability to hijack neuropilin-1”
As virologists the team had already noted that correlatively, this furin cleavage site is key to pathogenicity by looking at the sequences of past highly pathogenic viruses; however, the exact mechanism behind this was unknown.
When the sequence of the Spike protein of SARS-CoV-2 was discovered early in 2020, the Bristol researchers looked at the sequence of the Spike protein and noticed that it had a furin cleavage site that did not exist in the previous SARS-CoV. “This is a big difference,” Yamauchi noted. “The previous SARS-CoV is also fatal, but it was not spreading dramatically from human to human like SARS-CoV-2.”
Biochemical analysis
To understand this phenomenon in more detail, the team investigated the biochemical basis of the interaction between neuropilin-1 and the Spike protein. “We were lucky enough to have the live CoV-2 virus being handled in the same building,” said Yamauchi. This virus had been isolated from a British patient who was on the Diamond Princess cruise ship, had returned to the UK and then been hospitalised in Liverpool. “Not many researchers had access to the live virus at the university level, so we were very fortunate.”
Using biochemical techniques, the researchers focused in on the details of the interaction and found a three-dimensional binding pocket present on the surface of neuropilin-1 that the viral Spike protein engaged with, thanks to work from Brett Collins at the University of Queensland, Australia. “One role of neuropilin-1 in the cell’s context is to promote uptake of a ligand and disseminate that ligand deeper in the tissue,” said Yamauchi. “The SARS-CoV-2 Spike is a ligand for neuropilin-1 and therefore promotes the virus to enter cells and spread. Our hypothesis was that the reason this virus spreads quite dramatically between humans could be due to viral Spike’s ability to hijack neuropilin-1.” He noted that nasal epithelia express a lot of neuropilins and suggested perhaps this relates to the loss of taste, smell and neurological symptoms of SARS-CoV-2, citing a second study led by researchers in Munich and Helsinki.4 Such symptoms are absent from the previous SARS-CoV, so could be mediated by neuropilin-1.
 Following analysis of the structural confirmation of interactions between neuropilin-1 and the Spike protein, the researchers looked to target that interaction with small molecule inhibitors. “Because neuropilin-1 has been studied well in the context of cancer research, there are antagonists already commercially available,” said Yamauchi. Tambet Teesalu, from the University of Tartu, Estonia, a co-author in this study, also generated monoclonal antibodies (mAbs) to block the interaction between the Spike protein and neuropilin-1. These were able to significantly decrease the infectivity of the live virus compared to a control mAb targeting avian influenza A virus (H11N3) hemagglutinin, raising the possibility that antibodies or drugs targeting neuropilin-1 could be used to prevent the viral Spike protein from exploiting it to enter human cells.
Following analysis of the structural confirmation of interactions between neuropilin-1 and the Spike protein, the researchers looked to target that interaction with small molecule inhibitors. “Because neuropilin-1 has been studied well in the context of cancer research, there are antagonists already commercially available,” said Yamauchi. Tambet Teesalu, from the University of Tartu, Estonia, a co-author in this study, also generated monoclonal antibodies (mAbs) to block the interaction between the Spike protein and neuropilin-1. These were able to significantly decrease the infectivity of the live virus compared to a control mAb targeting avian influenza A virus (H11N3) hemagglutinin, raising the possibility that antibodies or drugs targeting neuropilin-1 could be used to prevent the viral Spike protein from exploiting it to enter human cells.
“There is definitely a role for neuropilin-1 in boosting the infectivity of SARS-CoV-2 and it is therefore undeniably a potential target for antiviral interventions,” concluded Yamauchi. He added that viruses are known to be very versatile and there is at least one other location where the SARS Spike protein binds to neuropilin-1. “We still have to fully understand the interaction interface of the Spike protein with neuropilin-1. With this knowledge, we may be able to target the interaction with stronger antiviral effects than what we have shown in this paper.”
Finally, Yamauchi clarified that neuropilin-1 is already being targeted by therapies, with programmes in lung inflammation control and certain cancers, some of which were in Phase II clinical trials in the US. “It is definitely a possibility that we could repurpose these already developed drugs for COVID-19 and future pandemics that hijack neuropilin-1 via the C-end rule motif. This is exciting science.”
Read the full paper…
To view the full paper, click here.
View the research…
To view a short video of the research, click here.
 Yohei Yamauchi is a virologist and ERC (European Research Council) investigator passionate about understanding the mechanism of viral infections. He is particularly known for his work on virus entry. Yohei – originally from Japan – trained as a medic and dermatologist in Nagoya before taking on a research career to study the interface of virus and host. He feels viruses are the best ‘cell biologists’ and investigating viruses can teach researchers a lot about the cells’ functions. Since 2016, he is Associate Professor at the University of Bristol and runs his own research group on virus cell biology with emphasis on the influenza A virus and now COVID-19.
Yohei Yamauchi is a virologist and ERC (European Research Council) investigator passionate about understanding the mechanism of viral infections. He is particularly known for his work on virus entry. Yohei – originally from Japan – trained as a medic and dermatologist in Nagoya before taking on a research career to study the interface of virus and host. He feels viruses are the best ‘cell biologists’ and investigating viruses can teach researchers a lot about the cells’ functions. Since 2016, he is Associate Professor at the University of Bristol and runs his own research group on virus cell biology with emphasis on the influenza A virus and now COVID-19.
Key collaborators
- Biochemistry: Peter Cullen, School of Biochemistry; Boris Simonetti, senior postdoc in the Cullen group (UoB).
- SARS-CoV-2 virology: Andrew Davidson and David Matthews, School of Cellular and Molecular Medicine (UoB); Julian Hiscox, University of Liverpool.
- Structural biology: Brett Collins, University of Queensland, Australia.
- C-end rule expert: Tambet Teesalu, University of Tartu, Estonia.
References
- Daly J, Simonetti B, Klein K, Chen K, Williamson M, AntónPlágaro C et al. Neuropilin-1 is a host factor for SARS-CoV-2 infection. Science. 2020;370(6518):861-865.
- Petersen E, Koopmans M, Go U, Hamer D, Petrosillo N, Castelli F et al. Comparing SARS-CoV-2 with SARS-CoV and influenza pandemics. The Lancet Infectious Diseases. 2020;20(9):e238-e244.
- Teesalu T, Sugahara K, Kotamraju V, Ruoslahti E. C-end rule peptides mediate neuropilin-1-dependent cell, vascular, and tissue penetration. Proceedings of the National Academy of Sciences. 2009;106(38):16157-16162.
- Cantuti-Castelvetri L, Ojha R, Pedro L, Djannatian M, Franz J, Kuivanen S et al. Neuropilin-1 facilitates SARS-CoV-2 cell entry and infectivity. Science. 2020; 370(6518):856-860.
Related topics
Disease Research, Drug Targets, Research & Development, Small Molecules, Structural Biology, Targets
Related conditions
avian influenza, Covid-19, Ebola, Middle East Respiratory Syndrome (MERS), Severe Acute Respiratory Syndrome (SARS)
Related organisations
BioNTech, Bristol University, Moderna, Pfizer, Queensland University, University of Tartu
Related people
Boris Simonetti, Brett Collins, Dr Yohei Yamauchi, Peter Cullen, Tambet Teesalu