Novel phage display technique could lead to new peptide-based therapeutics
Posted: 20 March 2020 | Hannah Balfour (Drug Target Review) | No comments yet
Researchers have shown that incorporating noncanonical amino acids (ncAAs) in phage display expands the range of peptides it can identify.
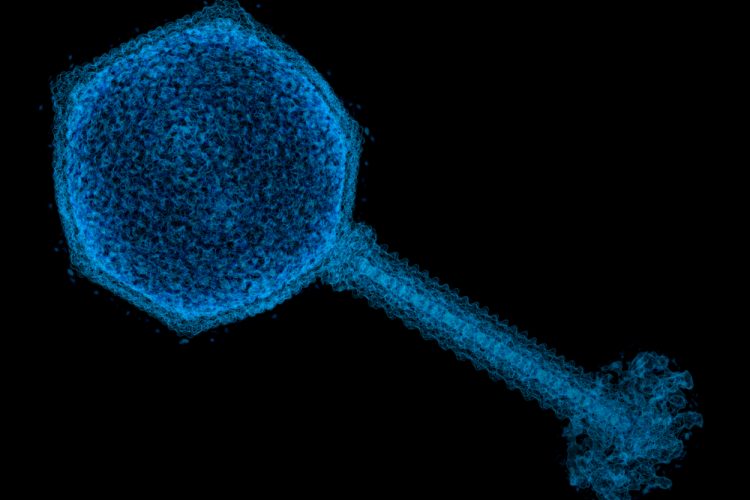

Researchers have developed a novel way of adding more complex peptides containing noncanonical amino acids (ncAAs) into viruses which would equip them with special properties, such as degradation resistance and targeted protein binding capabilities.
The research team, led by Texas A&M University, US, chemist Dr Wenshe Liu, identified that these properties could include that only viruses with ncAAs containing peptides could reproduce, enabling them to increase the ability of phage display to identify peptides for use as drugs. They suggest the use of ncAAs to expand the genetic code of bacteriophages could lead to new peptide-based therapeutics.
Published in Nature Communications, the paper shows how they incorporated ncAAs into phage displays and used them to identify novel peptides.
“Phage display uses viruses, or phages, to ‘fish out’ specific peptides from a pool of millions of different peptide variants; however, it is very difficult to use this technique to find peptides containing ncAAs,” added Dr Jeffery Tharp, a postdoctoral associate at Yale University, US, and lead author of the paper. “In our paper, we developed a new method of phage display that allows for easy retrieval of potential peptide drugs containing diverse ncAAs. In addition, we used our new technique to identify novel peptides containing ncAAs that are very strong inhibitors of sirtuin 2 – an enzyme that is involved in regulating human lifespan and is a promising drug target for the treatment of human cancers.”
Dr Tharp added: “This technique allows ncAAs with unique structures to be incorporated into the phage peptides, which can help identify more potent peptide drugs. In addition, we can include reactive ncAAs into the phage peptides, which can potentially be used to make better materials and drug delivery systems.”
The team stated they will continue to use the novel phage display technique to search for other peptides containing ncAAs that inhibit enzymes related to human disease and develop other methods that could expand the utility of phage displays.
Related topics
Analysis, Analytical Techniques, Bacteriophages, Drug Development, Drug Leads, Drug Targets, Research & Development, Screening, Technology, Therapeutics
Related organisations
Texas A&M University
Related people
Dr Wenshe R. Liu