Glycomimetics could prevent COVID-19 from infecting human cells
Posted: 21 April 2020 | Hannah Balfour (Drug Target Review) | No comments yet
Glycomimetics, a novel class of antivirals, prevented influenza, herpes viruses and papillomaviruses from infecting cells in animal models and could show similar efficacy against COVID-19.

A novel class of antiviral compounds could be effective in preventing COVID-19 infecting human cells, say researchers. Glycomimetics, a synthetic version of the glycosaminoglycans (GAGs) found on cell surfaces, showed efficacy in preventing influenza, herpes virus and papillomavirus infections in both cell cultures and an animal model.
In order to replicate and cause disease, viruses infect host cells, inserting their genome into the cell and hijacking endogenous cellular machinery to replicate. One strategy for combatting viral infections is to block viruses from binding with host cell surface molecules, preventing them from infecting these cells.
A type of cell surface molecule viruses often bind to is formed of sugars and is called a GAG. These structures are highly negatively charged and in the past have been added externally in experiments, reducing viral infections. However, natural polysaccharides have side effects associated with their biological function or impurities.
To capitalise on the advantages of GAGs, a research team led by Heinrich-Heine University, Germany, Professor Dr Laura Hartmann, from the Institute of Macromolecular Chemistry, produced artificial versions called glycomimetics. These comprise a long synthetic scaffold with side chains where small sugar molecules attached; both glycomimetics with shorter chains, up to 10 lateral sugars, known as ‘oligomers’ and long chains with up to 80 sugars, called ‘glycopolymers’, have been created. The negative charge was mimicked by adding sulfate groups to the sugars.
The antiviral properties of the glycomimetics was then tested in cell lines by Professor Dr Mario Schelhaas from the Institute of Cellular Virology at University Hospital Münster, Germany. They first tested the glycomimetics against papillomaviruses which can cause cervical cancer and discovered that dependent on the length of the sugar side chains, the antiviral effect was different.
The long chain synthetic molecules were more effective than the short chain ones because they prevented the virus from attaching to the cells, where the oligomer glycomimetics displayed antiviral activity after the attachment of the virus to the cell.
Professor Schelhaas said: “It is very likely that the long-chain molecules occupy the binding sites of the virus to the cell and thus block those sites. The short-chain molecules apparently do not block these sites. The next step is to test our hypothesis that these molecules prevent the redistribution of proteins in the virus particle so that the viruses cannot infect the cell.”
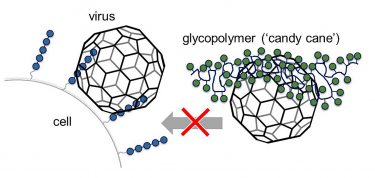

The artificially manufactured glycomimetics (green) attach to the viruses’ binding sites, which viruses normally use to attach to sugars (blue) on the cell surface [credit: L. Hartmann, M. Schelhaas].
The antiviral efficacy of the glycomimetics was also tested in animal models against papillomaviruses and four other viruses, including influenza and herpes viruses.
Professor Hartmann concluded: “Glycomimetics are thus promising compound molecules that could potentially be used in the fight against a large number of different viruses. The next thing to do is to examine the precise way in which the glycomimetics work and how they can be further optimised.”
Professor Schelhaas added: “Further research will focus on how fast viruses may adapt to this new class of compounds. With the short-chain molecules in particular, we are hopeful that viruses will find it harder to launch a counterattack.”
The paper was published in the Journal of American Chemical Society.
Related topics
Antimicrobials, Biopharmaceuticals, Cell Cultures, Disease Research, Drug Development, Drug Targets, Molecular Biology, Research & Development, Therapeutics
Related conditions
Coronavirus, Covid-19, Herpesvirus, Influenza, Papillomavirus
Related organisations
Heinrich-Heine University (HHU), Institute of Cellular Virology at University Hospital Münster
Related people
Professor Dr Laura Hartmann, Professor Dr Mario Schelhaas