Five recent CRISPR drug target discoveries
Posted: 21 November 2022 | Izzy Wood (Drug Target Review) | No comments yet
This article highlights five of the latest findings revealed using CRISPR that could be used in the development or design of new therapies.


Scientists and researchers investigating CRISPR-based techniques are finding promising therapeutic possibilities that with more research could be used to inform drug development. Here, we list five of the most recent target discoveries made using this technology.
1. SARM1: blocking nerve loss in neurodegenerative diseases
Researchers at Washington University School of Medicine, US, are working toward a treatment for neurodegenerative diseases, targeting SARM1, a key molecule involved in the death of axons.
The new studies, published in the Journal of Clinical Investigation,1,2 reveal surprising details about SARM triggering axon death, that underlies the development of neurodegenerative diseases. The research also points to novel therapeutic approaches for diseases defined by axon loss.
The study suggests that SARM1 contributes directly to axon loss and also plays a role in driving neuroinflammation that only serves to compound the problems”
Blocking this molecular executioner in a rodent model prevented axon loss, which has been implicated in many neurodegenerative diseases, from peripheral neuropathies to Parkinson’s disease and glaucoma to amyotrophic lateral sclerosis (ALS).
The researchers studied a rare progressive neuropathy syndrome. Sequencing patient genomes, they found that the axon loss was caused by genetic errors in the gene NMNAT2, whose normal function keeps SARM1 turned off.
Due to these genetic errors, SARM1 was constantly activated, triggering axon destruction. The researchers used the CRISPR gene-editing technique to reproduce these mutations in mice. These mice survived to adulthood but had worsening motor dysfunction, loss of peripheral axons and importantly, an infiltration of immune cells called macrophages.
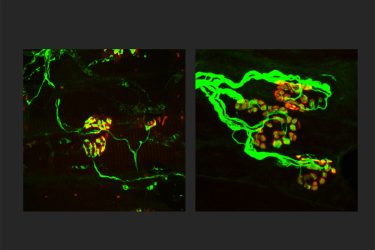

The researchers were surprised to find that reducing the number of macrophages reversed the axon loss and disease symptoms in the mice.
The study suggests that SARM1 contributes directly to axon loss and also plays a role in driving neuroinflammation that only serves to compound the problems. The findings also suggest that some neurodegenerative conditions could be treated with immune modulating drugs that block macrophages or other inflammatory immune cells.
“Our latest research suggests we also may be able to interfere with its ability to drive damaging neuroinflammation. We are hopeful this will lead to effective new therapies across a range of neurodegenerative and neuroinflammatory diseases.” said co-senior author Dr Jeffrey Milbrandt, the James S. McDonnell Professor and head of the Department of Genetics.
2. New treatment for CTLA-4 insufficiency
UCL, UK, researchers reveal a study that could lead to new treatments for CTLA-4 insufficiency, a rare disease of the white blood cells that normally help to control the immune system.3
In human cells, using the CRISPR-Cas system, the researchers targeted the faulty gene in T cells taken from patients with CTLA-4 insufficiency and repair the errors. CTLA-4 is a protein produced by T cells that helps to control the activity of the immune system.
This restored the levels of CTLA-4 in the cells to those seen in healthy T cells. They were also able to improve symptoms of the disease in mice with CTLA-4 insufficiency by giving them injections of gene-edited T cells.
Currently, the standard treatment for CTLA-4 insufficiency is a bone marrow transplant to replace the stem cells responsible for producing T cells.

Claire Booth, professor of gene therapy and paediatric immunology at UCL said: “Our approach has many positive aspects. By correcting the patient’s T cells, we think it can improve many of the symptoms of the disease, at the same time as being much less toxic than a bone marrow transplant. Collecting the T cells is easier and, correcting the T cells is easier.”
The gene editing approach uses the CRISPR-Cas9 system to target and snip the faulty CTLA-4 gene in two. Then a corrected sequence of DNA is delivered to the cell using a modified virus. This is then pasted over the faulty part of the gene using a cellular DNA repair mechanism known as homology-directed repair.
Booth concluded: “It is really exciting to think about taking this treatment forward to patients. If we can improve their symptoms and reduce their risk of getting lymphoproliferative disease this will be a major step forward.”
3. CRISPR-based microbiome editing
North Carolina State University (NC State), US, researchers show that viruses engineered with a CRISPR-Cas system can prevent bacterial defences and make selective changes to a targeted bacterium, even when other bacteria are in close proximity.4
Ultimately this study represents the next chapter of CRISPR delivery – using viruses to deliver CRISPR machinery in a complex environment”
The team deployed two different engineered bacteriophages to deliver CRISPR-Cas payloads for targeted editing of E. coli: first in a test tube and then within a synthetic soil environment created to mimic soil.
Both the engineered bacteriophages – T7 and lambda – successfully found and then delivered payloads to the E. coli host on the lab bench. These payloads expressed bacterial fluorescent genes and manipulated the bacterium’s resistance to an antibiotic. The team used lambda to deliver a so-called cytosine base editor to the E. coli host.
Professor Todd Klaenhammer, Bioprocessing and Nutrition Sciences at NC State, added: “We use a bacterial virus, a bacteriophage, to deliver CRISPR to bacteria, which is ironic because bacteria normally use CRISPR to kill viruses.”
“We used a base editor here as a kind of programmable on-off switch for genes in E. coli. Using a system like this, we can make highly precise single-letter changes to the genome without the double-strand DNA breakage commonly associated with CRISPR-Cas targeting,” said Matthew Nethery, a former NC State PhD student and lead author of the study.
“We see this as a mechanism to aid the microbiome. We can make a change to a particular bacterium and the rest of the microbiome remains unscathed,” Raldolphe Barrangou said. “This is a proof-of-concept that could be employed in any complex microbial community, which could translate into better plant health and better gastrointestinal tract health.
“Ultimately this study represents the next chapter of CRISPR delivery – using viruses to deliver CRISPR machinery in a complex environment.”
4. Novel UBA-inhibiting drug: TAK-243
Researchers at Virginia Commonwealth University (VCU) Massey Cancer Centre, US, have set their sights on a new therapeutic target for triple negative breast cancer (TNBC) with limited treatment options.
Through a CRISPR-Cas9 screening, the scientists were able to identify a specific enzyme called UBA1 that revealed itself as an ideal therapeutic target.
By using a novel UBA-inhibiting drug called TAK-243, they blocked the cellular function of UBA1 and effectively killed cancer cells in patient-derived breast tumours in mice.
This study, published in PNAS Nexus,5 is the first to suggest that UBA1 inhibitors could be effective in TNBC. TAK-243 has been tested in early phase trials, paving the way for potential testing in TNBC patients.
“We found that the majority of TNBC cells were uniformly susceptible to the antitumor effects of TAK-243,” said Dr Jennifer Koblinski, director of the Cancer Mouse Models Core and member of the Cancer Biology research programme at VCU. “In addition to demonstrating this drug’s success at the local site of the primary breast cancer, our findings demonstrate that TAK-243 can also shrink tumours in various organs after the disease has spread.”
The researchers also determined that the c-MYC gene, an important and difficult drug target in TNBC, can be harnessed to co-operate with TAK-243 to initiate a cellular stress response and enhance the drug’s ability to fight TNBC.
“The implementation of targeted therapies — drugs that target a specific genetic defect — has been revolutionary in treating different cancers, including breast cancer,” added Dr Anthony Faber, co-leader of the Developmental Therapeutics research programme in Cancer Research at VCU. “Genomic and clinical evidence suggests that the implementation of targeted therapies in the treatment of TNBC will require an expansion of potential targets. Our study may have identified a key and novel target for the development of new therapies.”
The next steps of this research include exploring the use of TAK-243 in TNBC with the drug company developing it and evaluating other targets in the UBA1 pathway that also demonstrate efficacy.
5. SOX9 facilitates cancer relapse
Researchers at Uppsala University, Sweden, have discovered that the protein: SOX9 makes tumour cells rest and become insensitive to radiation treatment.6
In the study, the team has showed that certain cells within the tumour bulk did not divide as often as other cancer cells did, which made them less sensitive to irradiation.
They also found an accumulation of a specific protein, SOX9, in recurrent samples from patients that had been operated before and after they developed a relapse.
“In experiments where we knocked out SOX9 with the CRISPR-Cas9 genetic scissors, tumour cells lost their capability to relapse, which suggests that SOX9 is important for this process,” said Fredrik Swartling, lead author.
The researchers further examined how substances that inhibited SOX9 influenced the development of relapses in animal models. Using bioinformatic analyses they discovered a few drugs that are used for other treatments that unexpectedly had a suppressing effect on SOX9 in relapses.
“We hope that our discovery could lead to more specific treatments against those SOX9-positive, slow dividing cancer cells. Eventually, it might improve the possibilities to treat children with medulloblastoma who have the highest risk of developing relapses,” said Anna Borgenvik, a postdoc in the research group.
The research group hopes that the results could eventually lead to better treatments for children that have the highest risk to develop relapses.
References
1. Yamada Y, Strickland A, Sasaki Y, et al. A SARM1/mitochondrial feedback loop drives neuropathogenesis in a Charcot-marie-tooth disease type 2A RAT model. Journal of Clinical Investigation. 2022;
2.Dingwall CB, Strickland A, Yum SW, et al. Macrophage depletion blocks congenital SARM1-dependent neuropathy. Journal of Clinical Investigation. 2022;
3.Fox TA, Houghton BC, Petersone L, et al. Therapeutic gene editing of T cells to correct CTLA-4 insufficiency. Science Translational Medicine. 2022;14(668).
4.Nethery MA, Hidalgo-Cantabrana C, Roberts A, Barrangou R. CRISPR-based engineering of phages for in situ bacterial base editing. Proceedings of the National Academy of Sciences. 2022;119(46).
5.Jacob S, Turner TH, Cai J, et al. Genomic screening reveals ubiquitin-like modifier activating enzyme 1 as a potent and druggable target in c-myc-high triple negative breast cancer models. PNAS Nexus. 2022;1(5).
6. Borgenvik A, Holmberg KO, Bolin S, et al. Dormant sox9-positive cells facilitate Myc-driven recurrence of Medulloblastoma. Cancer Research. 2022;
Related topics
Animal Models, CRISPR, DNA, Drug Development, Drug Discovery, Drug Discovery Processes, Genomics, Immuno-oncology, Immunology, Immunotherapy, Sequencing, Targets, Therapeutics
Related conditions
CTLA-4 insufficiency, Neurodegenerative diseases, triple negative breast cancer (TNBC)