Finding drugs for breast cancer targets
Posted: 7 March 2022 | Ria Kakkad (Drug Target Review) | No comments yet
A scientist at the University of Houston receives a $2 million grant to innovate computer-aided drug discovery for breast cancer.
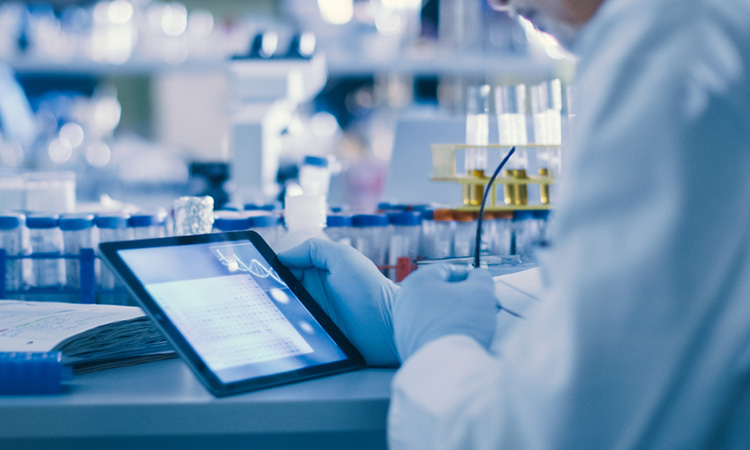


Gül Zerze, assistant professor at the University of Houston, US, is one of 12 researchers to receive a $2 million grant from the Cancer Prevention and Research Institute of Texas, US. With the grant, Zerze is setting up a laboratory to develop therapeutics that will focus on traditionally undruggable targets in cancer, specifically breast cancer.
“One out of nearly six Texas women diagnosed with breast cancer will die of the disease. Importantly, Texan women of colour are disproportionately impacted by the high mortality rate compared to white Texan women (41 percent higher mortality rate reported for Black Texan women in 2016). This high mortality rate, despite the substantial efforts made for early diagnosis, calls for better therapeutics urgently,” said Zerze.
Approximately 70 percent of proteins implicated in human cancers are either intrinsically disordered proteins (IDPs) or have large intrinsically disordered regions, and many of these targets are considered ‘undruggable’ due to the scarcity of high-resolution methods that can offer a fundamental understanding of them.
“Computational and data science methodologies offer a promising avenue to fill in this gap to enable developing drugs against these traditionally undruggable targets,” said Zerze, whose methodology will include rapid screening.


Gül Zerze, assistant professor in the William A. Brookshire Department of Chemical and Biomolecular Engineering at the University of Houston, is targeting formerly undruggable targets in breast cancer.
[Credit: University of Houston].
While there has been significant progress in cancer treatment over the past two decades, many cancer targets have yet to be drugged. Among promising therapeutics are transcription factors (TFs), which are proteins involved in converting (or transcribing) DNA into RNA. TFs contain large amounts of disordered proteins, which participate in transcriptional condensates that form via liquid-like phase separation (LLPS).
“Transcriptional condensates are shown to be aberrant in tumour cells, but the progress to develop drugs against TFs that participate in LLPS has been limited by the extremely dynamic nature of activation domains of TFs. We are developing a computational platform that will enable discovering drugs against these aberrant condensates by systematically interrogating the way transcription factors form, through the liquid-like phase separation of intrinsically disordered regions,” explained Zerze.
The CPRIT project hopes to promote commercialisation in Texas and founding companies to contribute to the local economy.
Related topics
Disease Research, DNA, Drug Development, Drug Discovery Processes, Funding, Protein, Research & Development, Technology
Related conditions
Breast cancer
Related organisations
University of Houston
Related people
Gül Zerze