Researchers develop ACE2 decoy to neutralise SARS-CoV-2
Posted: 10 November 2020 | Hannah Balfour (Drug Target Review) | No comments yet
Using their de novo protein design strategy, researchers engineered human angiotensin converting enzyme 2 (hACE2) protein decoys that can protect cells from SARS-CoV-2 infection.
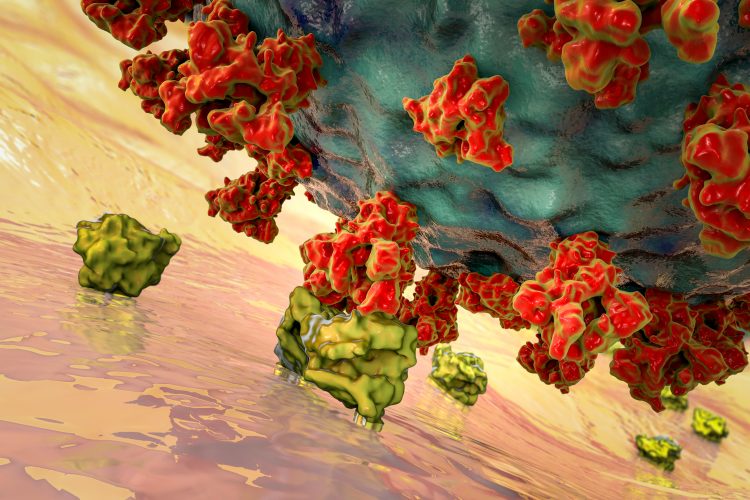

To infect cells within the respiratory tract, the Spike (S) protein on the surface of SARS-CoV-2 must bind to a receptor on human cells called angiotensin converting enzyme 2 (ACE2). The receptor binding domains (RBDs) are the section of the S protein which binds to hACE2. Disrupting this interaction has become the focus of a lot of research, as it would prevent infection; however, when neutralising antibodies have been developed to target the S protein, it has been shown to evade them by mutating.
To address this issue, Thomas Linsky and colleagues created a computational protein design strategy to enable the development of therapeutics that may be more resistant to SARS-CoV-2 mutational escape. In their study published in Science, the team describe the design, validation and optimisation of stable hACE2 protein decoys which mimic the protein receptor interface to which SARS-CoV-2 binds.
They developed roughly 35,000 computational decoys and took the top-ranking ones into further testing. From these experiments they identified CTC-445.2 as the best decoy. According to the study, CTC-445.2 binds with low nanomolar affinity and high specificity to the RBD of the S protein. Using cryogenic electron microscopy (cryo-EM) they observed that CTC-445.2 also simultaneously bind to all three RBDs.
The team also developed a bivalent decoy based on CTC-445.2, that they called CTC-445.2d. According to the researchers, with the bivalent decoy there was approximately a 10-fold improvement in binding. CTC-445.2d potently neutralised SARS-CoV-2 infection in vitro and when a single intranasal prophylactic dose was administered to a Syrian hamster model 12 hours before viral challenge, it allowed all animals to survive the lethal dose with only modest weight loss.
The team concluded that because the decoy replicates the S protein target interface, it is intrinsically resilient to viral mutational escape.
Related topics
Analytical Techniques, Bioengineering, Drug Development, Drug Leads, Imaging, In Vitro, In Vivo, Protein, Proteomics, Research & Development, Therapeutics
Related conditions
Coronavirus, Covid-19
Related people
Thomas Linsky