Five developments in oncology targets
Posted: 30 July 2020 | Hannah Balfour (Drug Target Review) | No comments yet
Drug Target Review explores some of the newest oncologic drug targets, including those for glioblastoma, lung cancer and breast cancer.
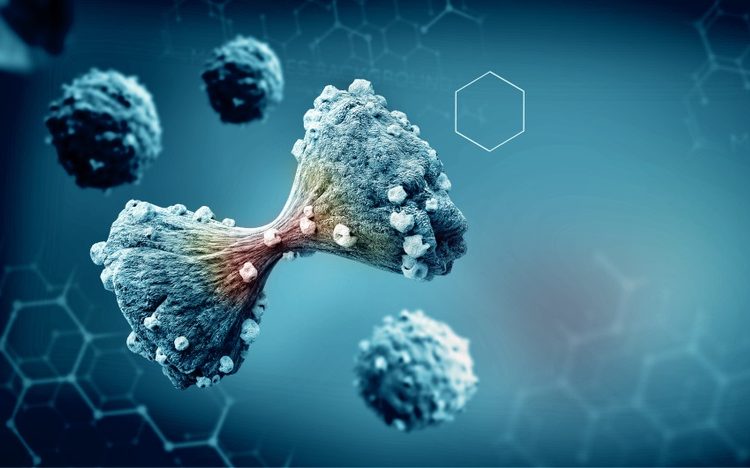

Cross-linking protein implicated in cancer is vital to cell division
According to new research published in Developmental Cell, the PRC1 protein is an essential component in cell division. Acting as a viscous glue, the protein creates resistance as cells pull chromatids to either end of the single dividing cell.
The team explains that this process controls the speed that DNA is separated into daughter cells, suggesting that the over-abundance of PRC1 characteristic observed in many cancer types (including prostate, ovarian and breast cancers) may be causing the cells to divide unevenly, resulting in chromosomal abnormalities and driving cancer growth and mutation.
“PRC1 produces a viscous frictional force, a drag that increases with speed,” said Scott Forth, an Assistant Professor of Biological Sciences and member of the Center for Biotechnology and Interdisciplinary Studies at Rensselaer Polytechnic Institute in the US. “The friction it produces is similar to that of water – if you try to move your hand through water slowly, you move easily, but if you push your hand fast, the water pushes back hard.”
The team explained that correct cellular division relies on physical forces produced by motor proteins and microtubules, essentially ripping pairs of chromatids apart so a single cell of the cellular DNA ends up in each daughter cell. The mitotic spindle is an element of cellular machinery that uses mechanical forces – push, pull and resistance – to complete the task.
“We think the force PRC1 produces is integrating and dampening out cellular motions as the DNA is separated so that ultimately, you get the correct rate of chromosome segregation,” Forth said. However, if this process goes awry due to too much or too little PRC1, it can cause uncontrollable cancer growth.
Expansion stress drives breast cancer aggressiveness, finds study
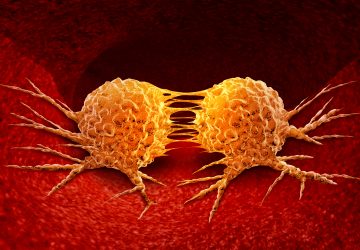

Expansion stress is a condition often seen in solid cancer tumours when, as they grow, biomechanical forces in the tumour microenvironment cause compression within the tumour, tension at its periphery and altered interstitial fluid flow.
The interdisciplinary team from the University of Alabama at Birmingham (UAB), US, also suggested that the biomechanical forces may modulate the immune response through cancer cell-immune cell crosstalk.
To explore how these biomechanical forces may be impacting breast cancer growth, the researchers created a tissue-engineered, three-dimensional (3D) breast cancer mimetic system that recapitulates the in vivo growth of breast cancer cells. It includes tumour-associated fibroblasts, endothelial cells and immune cells, within a physiologically relevant extracellular matrix.
According to the team, the biomechanical forces significantly altered the proteome of breast cancer cells and enhanced exosome production. These tumour cell-secreted exosomes are one of the intercellular mediators of signalling in the tumour microenvironment and are now considered key regulators of tumour progression.
In the study, the exosomes promoted aggressive tumour cell growth, induced immune suppression and altered immune cell polarisation in the tumour microenvironment. Furthermore, the researchers recently engineered an oscillatory compression device for real-time application of biomechanical force on orthotopic mammary tumours in mice, which allowed them to observe exosome-mediated immunosuppression and aggressive tumour growth.
The researchers concluded that their study suggests that exposure to mechanical strain promotes invasive and pro-tumourigenic phenotypes of breast cancer cells. They also added that mechanical strain also impacted the growth and proliferation of cancer cells, altered exosome production and induced immunosuppression in the tumour microenvironment.
The paper was published in Laboratory Investigation.
Exploring the causes of lung cancer in non-smokers could improve treatment strategies
Researchers studying lung cancers revealed that the cancers observed in non-smokers are distinctly different to those in smokers, with different genetic underpinnings dependent on age, and lifestyle. They suggest that some patient populations may respond to targeted treatments better than others.
The study, conducted in Taiwan, was co-led by scientists at The Institute of Cancer Research.
According to the scientists, around 10-15 percent of lung cancers in the UK occur in people who have never smoked, but in East Asia this proportion is much higher, especially among women.
The study, published in Cell, details the analyses of tumour samples from 103 lung cancer patients from Taiwan, the majority of which were non-smokers. The researchers conducted a detailed analysis of genetic changes, gene activation, protein activity and cellular ‘switches’ in lung cancer to develop a comprehensive picture of the biology of lung cancers in non-smokers.
Looking at the genetics and the related proteins produced by cancer cells in the tumour samples, the team found that some early-stage lung tumours in non-smokers were biologically similar to more advanced disease in smokers.
Tumours in females typically had a defect in the epidermal growth factor receptor (EGFR) gene, whereas in men the most common mutations were in the KRAS (Ki-ras2 Kirsten rat sarcoma viral oncogene homologue) and adenomatous polyposis coli (APC) genes. The team suggested this could cause the two sexes to respond differently to treatments.
The study also identified a pattern of genetic changes involving the APOBEC (apolipoprotein B mRNA editing catalytic polypeptide-like) gene family in 75 percent of tumours in female patients under the age of 60 and 100 percent of with no mutation in the EGFR gene.
APOBEC proteins play an important role in the function of the immune system but can be hijacked by cancers to speed up their evolution, which can expedite the development of drug resistance. According to the team, patients without EGFR defects tend to respond better to immunotherapy and suggest that testing for APOBEC could help identify women more likely to respond.
Finally, the team identified 65 proteins that were overactive in the tumours that could be targeted by existing drug candidates.
While the new study included patients treated in Taiwan, the researchers believe that many of their findings could be applicable to UK patients. They will be validating their findings in larger studies and beyond Asia.
Dr Jyoti Choudhary, Team Leader in Functional Proteomics at ICR, UK, said: “We carried out the most comprehensive study ever conducted into the biology of lung cancers in an East-Asian population with a high proportion of non-smokers and found that their disease is molecularly diverse and distinct from what we classically see in smokers.

Dr Emily Armstrong, Research Information Manager at Cancer Research UK, said: “In order to beat cancer, we need to understand all the ways it can develop… Understanding the difference between lung cancers in smokers and non-smokers could be vital for providing patients with the most appropriate treatment.”
Glioblastoma stem cells drive tumour growth, study finds
Scientists have identified a population of cells within glioblastomas from which all other cancerous cells within the tumour arise. They suggested that targeting these ‘glioblastoma stem cells (GSCs)’ could be a potent future treatment option.
Tumour heterogeneity, or the genetic variation of tumour cells, is one reason why brain cancers can be resistant to treatments. After a treatment, resistant cells remain and subsequently repopulate the tumour.
In a new study, researchers identified a cancer cell hierarchy, indicating that each cell originates from a single cancer cell type, coined glioblastoma stem cells by the team. They suggested that targeting these cells with drug interventions could therefore slow cancer growth.
The team began by sequencing the RNA from 55,000 glioblastoma cells and 20,000 normal brain cells. They identified five main cancer cell types within each tumour and found that these are similar to cell types found within the human brain.
One cell type described was a progenitor GSC, a cell type from which all other glioblastoma cells would develop. They further showed that there was a cellular hierarchical organisation to the cancer that originates from progenitor GSCs.
According to the team, GSCs divided at a faster rate than the mature cancer cells. In fact, they made up most of the dividing cells within the tumour, despite making up a relatively small proportion of the cells within the whole tumour. These rapidly dividing cells are the earliest detectable cancer cells in the hierarchy, suggesting they are a promising target for therapy.
The researchers identified that molecular vulnerabilities of the progenitor GSCs and targeted them in experiments. They found that progenitor GSC survival and proliferation decreased as a result of these interventions and that in pre-clinical disease models, this reduced tumour growth and increased survival of the animals.
“Our work has gone a long way to resolve the complexity of glioblastoma heterogeneity and provides a new framework to reconsider the nature of glioblastoma,” explained research leader, Dr Kevin Petrecca, a neurosurgeon and brain cancer researcher at The Neuro (Montreal Neurological Institute and Hospital) of McGill University, part of the McGill University Health Centre in Canada. “As part of this work, our study also shows, in contrast to decades long dogma, that glioblastoma stem cells are the most rapidly dividing cancer cells in the tumour, and we identified new ways to target these cells. There is still much work to be done. Understanding how these cancer cells interact with the cancer microenvironment is not well understood in this disease, but this study serves as a good starting point to begin to understand how glioblastoma originates and evolves prior to treatments.”
This study was published in Nature Communications.
AVIL gene: a potential drug target for glioblastoma
Scientists have identified AVIL as an oncogene responsible for glioblastoma, the deadliest brain tumour. According to the researchers, the discovery offers a promising new treatment target for this cancer, as this gene is vital to the survival of glioblastoma cancer cells.
“Glioblastoma is one of the most deadly cancers. Unfortunately, there is no effective treatment option for the disease,” said researcher Dr Hui Li, of the University of Virginia School of Medicine and the UVA Cancer Center’s Department of Pathology, both US. “The novel oncogene we discovered promises to be an Achilles’ heel of glioblastoma, with its specific targeting potentially an effective approach for the treatment of the disease.”

In their experiments, blocking AVIL destroyed glioblastoma cells within mice, but had no effect on healthy tissue. The team concluded that therapies targeting this gene are likely to be highly effective and selective.
“AVIL is overexpressed in 100 percent of glioblastoma cells and clinical samples, and is expressed at even higher levels in so-called ‘glioblastoma stem cells’, but hardly expressed in normal cells and tissues,” said Li. “Silencing the gene wiped out glioblastoma cells in culture and prevented animal xenografts, while having no effect on normal control cells. Clinically, high AVIL expression correlates with worse patient outcomes. These findings and classic transformation assays proved AVIL being a bona fide oncogene.”
The researchers said that the AVIL gene plays a “critical role” in glioblastoma development and survival. Li added that the strategy they used to identify this oncogene, studying a structural variant identified in a paediatric cancer, could be used to identify other genes driving other adult cancers.
The paper was published in Nature Communications.
Related topics
Disease Research, Drug Targets, Oncology, Research & Development
Related conditions
Breast cancer, Cancer, Glioblastoma
Related organisations
Cancer Research UK, McGill University, Rensselaer Polytechnic Institute, The Institute of Cancer Research, University of Alabama at Birmingham, University of Virginia School of Medicine
Related people
Dr Emily Armstrong, Dr Hui Li, Dr Jyoti Choudhary, Dr Kevin Petrecca, Scott Forth