Researchers simulate hepatitis B infection using computational method
Posted: 8 September 2021 | Anna Begley (Drug Target Review) | No comments yet
The new study modelled the process of capsid disassembly of the hepatitis B virus at an atomic level to help develop targeted therapies.
A team at the Beckman Institute, US, have used new computational techniques to model the process of capsid disassembly of the hepatitis B virus (HBV) at an unprecedented atomic level, potentially leading to targeted therapies against the virus in the future.
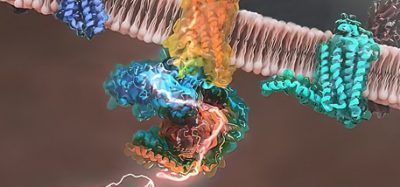
The hepatitis B capsid is an icosahedral structure about 36 nanometres in diameter and composed of 240 identical proteins. Capsid disassembly allows the virus to release its own genetic material into the host cell’s nucleus and eventually use the host cell’s replication machinery to multiply. According to the team, understanding this process on an atomic level is imperative to combat HBV and other similar infections.
The researchers applied atomistic molecular dynamics simulation to an entire HBV capsid subjected to a uniform mechanical stress protocol. “Over the past few decades, the developments of advanced simulation software such as NAMD… and a more accurate treatment of interactions between atoms, enabled us to simulate a system of this size and complexity,” explained lead author Dr Zhaleh Ghaemi.
The team then identified which areas of the hepatitis B capsid impact the disassembly process. Surprisingly, specific regions of the capsid protein were found to contribute to the breakage more than others and not in the pattern that had previously been surmised based on mechanical properties alone.
By probing and investigating capsid disassembly, which was set to efficiently occur in just a few nanoseconds of simulation, the team predicted that the first cracks that lead to disassembly are a result of the capsid structure expanding by just 2.5 percent. “The work here is based on simulations, but we can confirm the simulations with experiments that involve mutating specific amino acids predicted to be ‘hot spots’ for disassembly,” Ghaemi stated.
The researchers hope that their findings will not only advance the study of HBV to find new targeted therapies, but also that their technique will further understandings in other scientific fields. “Tools like this equip us with a computational microscope by which we can watch complex molecular motions and phenomena at an extremely high resolution that cannot be yet achieved otherwise,” said Professor Emad Tajkhorshid.
The study was published in PNAS.
Related topics
Drug Targets, Informatics, Molecular Biology, Molecular Modelling, Molecular Targets, Nanoparticles, Protein, Small molecule, Structural Biology, Target Molecule, Technology
Related conditions
Hepatitis B
Related organisations
Beckman Institute
Related people
Emad Tajkhorshid, Zhaleh Ghaemi