Virtual reality could be used to design new drugs, say researchers
Posted: 12 March 2020 | Victoria Rees (Drug Target Review) | No comments yet
Researchers have used virtual reality (VR) to control how drugs bind to their protein targets, which they say could be useful for designing new treatments.

Researchers report that they have used virtual reality (VR) as new way to design drugs, as well as understand how common medications work on a molecular level.
The study, conducted at the University of Bristol, UK, allowed the researchers to use VR and ‘step inside’ proteins to manipulate them and the drugs binding to them in atomic detail, using interactive molecular dynamics simulations in VR (iMD-VR). Employing this iMD-VR approach, researchers ‘docked’ drug molecules into proteins and were able to predict accurately how the drugs bind.
Professor Adrian Mulholland, from the University of Bristol’s Centre for Computational Chemistry and co-lead of the work, said: “Many drugs work by binding to proteins and stopping them working. For example, by binding to a particular virus protein, a drug can stop the virus from reproducing. To bind well, a small molecule drug needs to fit snugly in the protein. An important part of drug discovery is finding small molecules that bind tightly to specific proteins and understanding what makes them bind tightly, which helps to design better drugs. To design new therapies, researchers need to understand how drug molecules fit into their biological targets. To do this, we use VR to represent them as fully three-dimensional (3D) objects. Users can then fit a drug within the ‘keyhole’ of a protein binding site to discover how they fit together.”
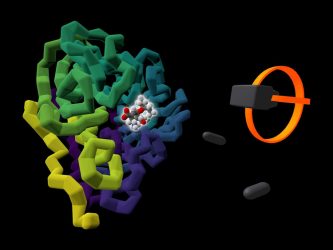
The commercially available influenza drug Tamiflu (highlighted by a glow) is shown bound to the viral protein it targets, neuraminidase (coloured in purple to yellow). A rendering of a user’s VR headset and controllers is presented alongside the protein (credit: University of Bristol).
In the study, the researchers attempted to bind drugs to protein targets such as influenza neuraminidase and HIV protease. The tests showed that they were able to predict correctly how the drugs bind to their targets. By pulling the drug into the protein, the scientists could build structures very similar to the drug complexes found in experiments.
The researchers also demonstrated that it is possible to unbind and rebind drugs from protein targets on a simulation timescale significantly shorter than the timescale of similar events observed using non-interactive molecular dynamics engines. The full unbinding and rebinding of the drug and proteins generated using iMD-VR were achieved by the researchers in less than five minutes of real time.
According to the researchers, their study shows how VR can be used effectively in structure-based drug design. It uses readily available VR equipment and an open source software framework.
Professor Mulholland added: “An important aspect of the work is that the drugs and their protein targets, are fully flexible; we model their structural changes and dynamics and users can manipulate them interactively to find how drugs interact with their biological targets. This is a really exciting and powerful way to model drug binding. We have shown in this work that it gives accurate results. These tools will be useful in the design and development of new drugs.”
The findings were published in PLOS ONE.
Related topics
Disease Research, Drug Targets, Imaging, Informatics, Protein, Research & Development, Robotics, Target Molecule, Technology
Related organisations
Bristol University
Related people
Professor Adrian Mulholland